Selected Publications
You can find a full publication list at Google Scholar.
Underline: trainees from Mostafavi lab; *: co-first authorship; $: co-senior authorship.
Preprints
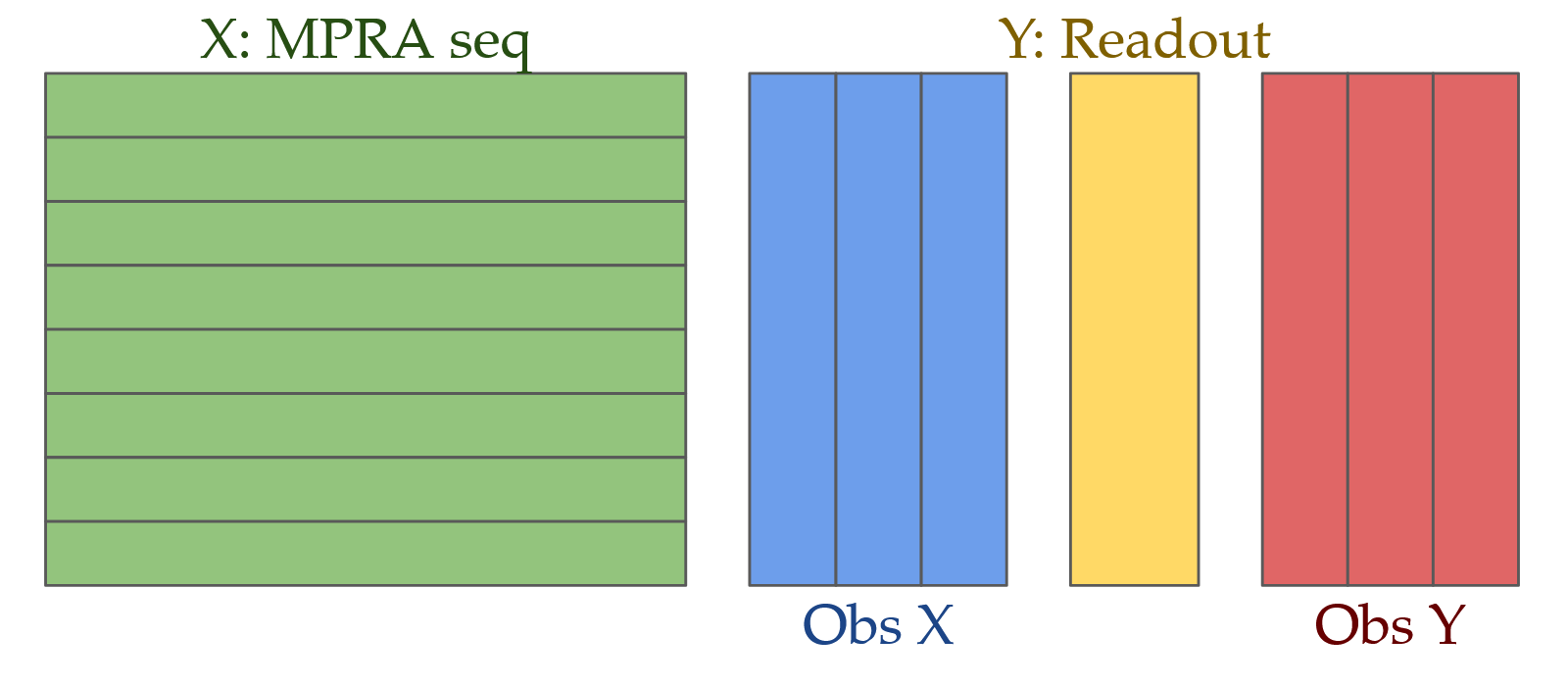
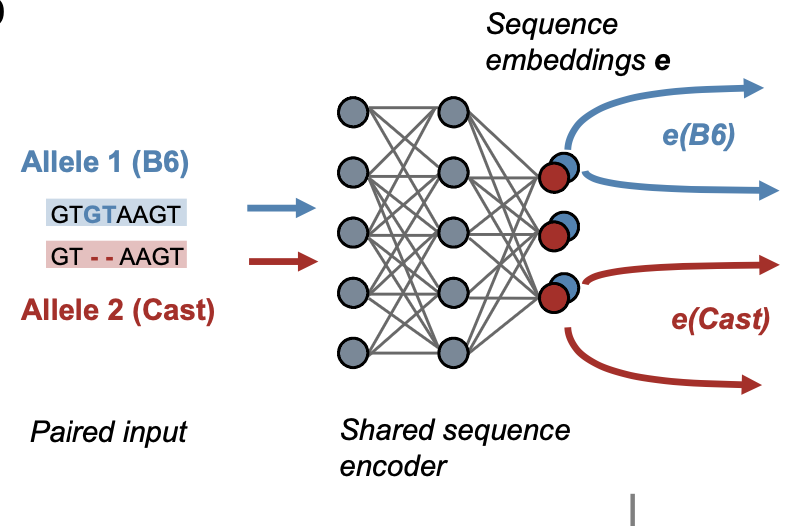
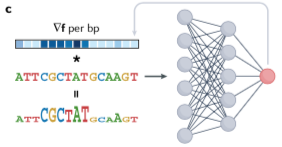
Deep genomic models of allele-specific measurements
Xinming Tu*, Alexander Sasse*,Kaitavjeet Chowdhary*,Anna Spiro, Liang Yang, Maria Chikina, Christophe Benoist,Sara Mostafavi. bioRxiv
2025
Investigating Data Size, Sequence Diversity, and Model Complexity in MPRA-based Sequence-to-Function Prediction
Yilun Sheng, Xinming Tu, Sara Mostafavi. bioRxiv
2025
Refining the cis-regulatory grammar learned by sequence-to-activity models by increasing model resolution
Nuria Chandra, Yan Hu, Jason D. Buenrostro, Sara Mostafavi$,Alexander Sasse$. bioRxiv
2025
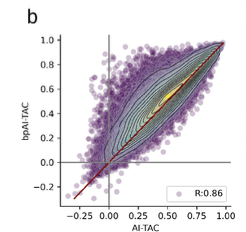
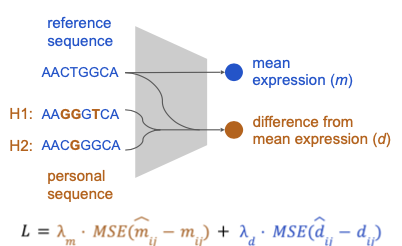
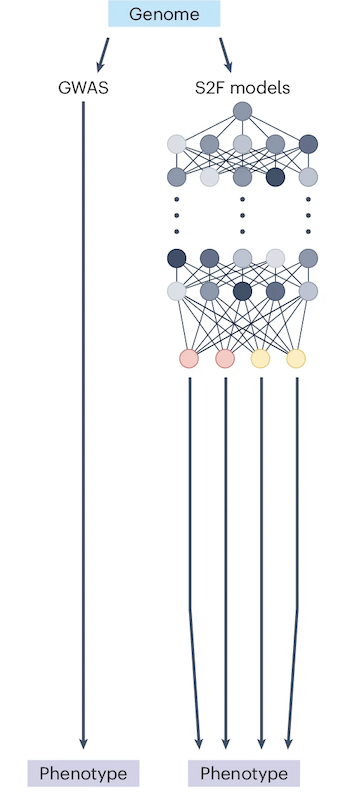
A scalable approach to investigating sequence-to-expression prediction from personal genomes
Anna Spiro*, Xinming Tu*, Yilun Sheng, Alexander Sasse, Rezwan Hosseini, Maria Chikina, Sara Mostafavi$. bioRxiv
2025
Selected Published Articles
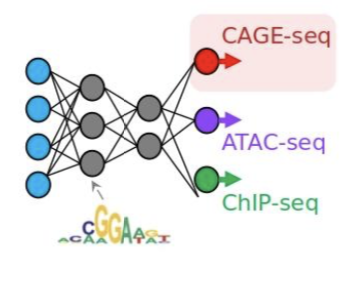
Unlocking gene regulation with sequence-to-function models
Alex Sasse, Maria Chikina$, Sara Mostafavi$. Nature Methods (Comment)
2024
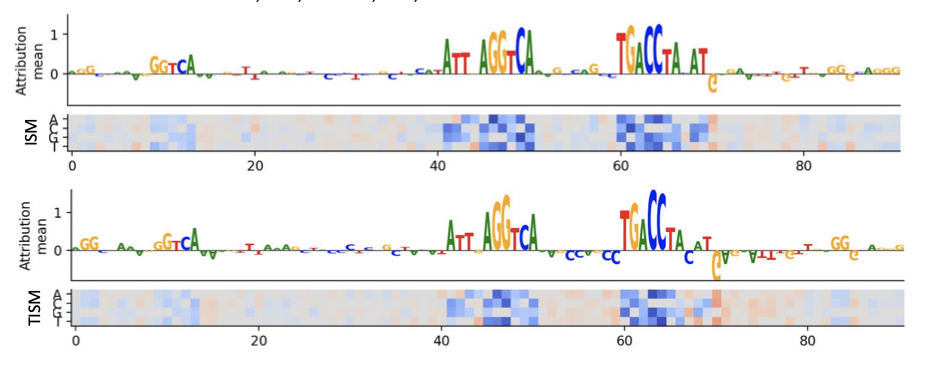
Quick and effective approximation of in silico saturation mutagenesis experiments with first-order Taylor expansion
Alex Sasse, Maria Chikina$, Sara Mostafavi$. RECOMB-seq
2024
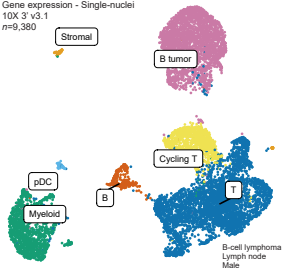
AIRE relies on Z-DNA to flag gene targets for thymic T cell tolerization
Yuan Fang, Kushagra Bansal, Sara Mostafavi, Christophe Benoist, Diane Mathis. Nature
2024
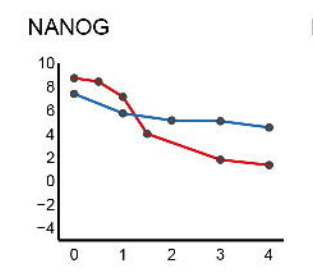
Benchmarking of deep neural networks for predicting personal gene expression from DNA sequence highlights shortcomings
Alex Sasse*, Bernard Ng*, Anna Spiro*,Shinya Tasaki, David Bennett, Chris Gaiteri, Phil De Jager, Maria Chinkina$, Sara Mostafavi$. Nature Genetics
2023
Cell sub-type specific effects of genetic variation in the aging and Alzheimer cortex
Masashi Fujita, Zongmei Gao, Lu Zeng, ..., Sara Mostafavi, David Bennett, Vilas Menon, Philip De Jager. Nature Genetics (Accepted)
2023
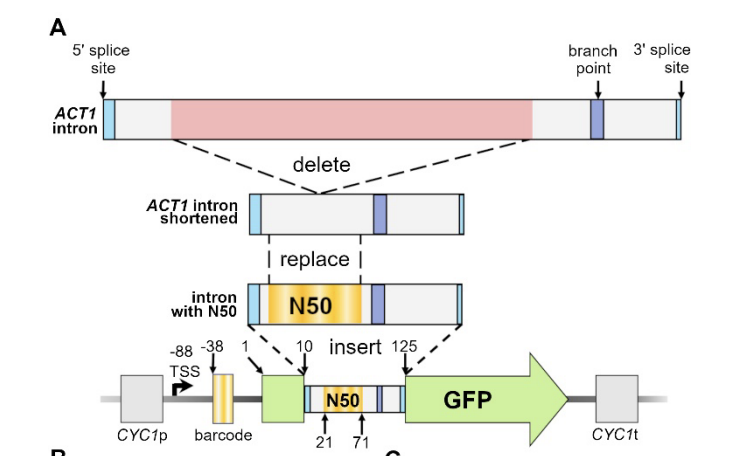
Impact of random 50-base sequences inserted into an intron on splicing in Saccharomyces cerevisiae
Molly Perchlik, Alex Sasse, Sara Mostafavi, Stan Fields, Josh T. Cuperus. RNA
2023
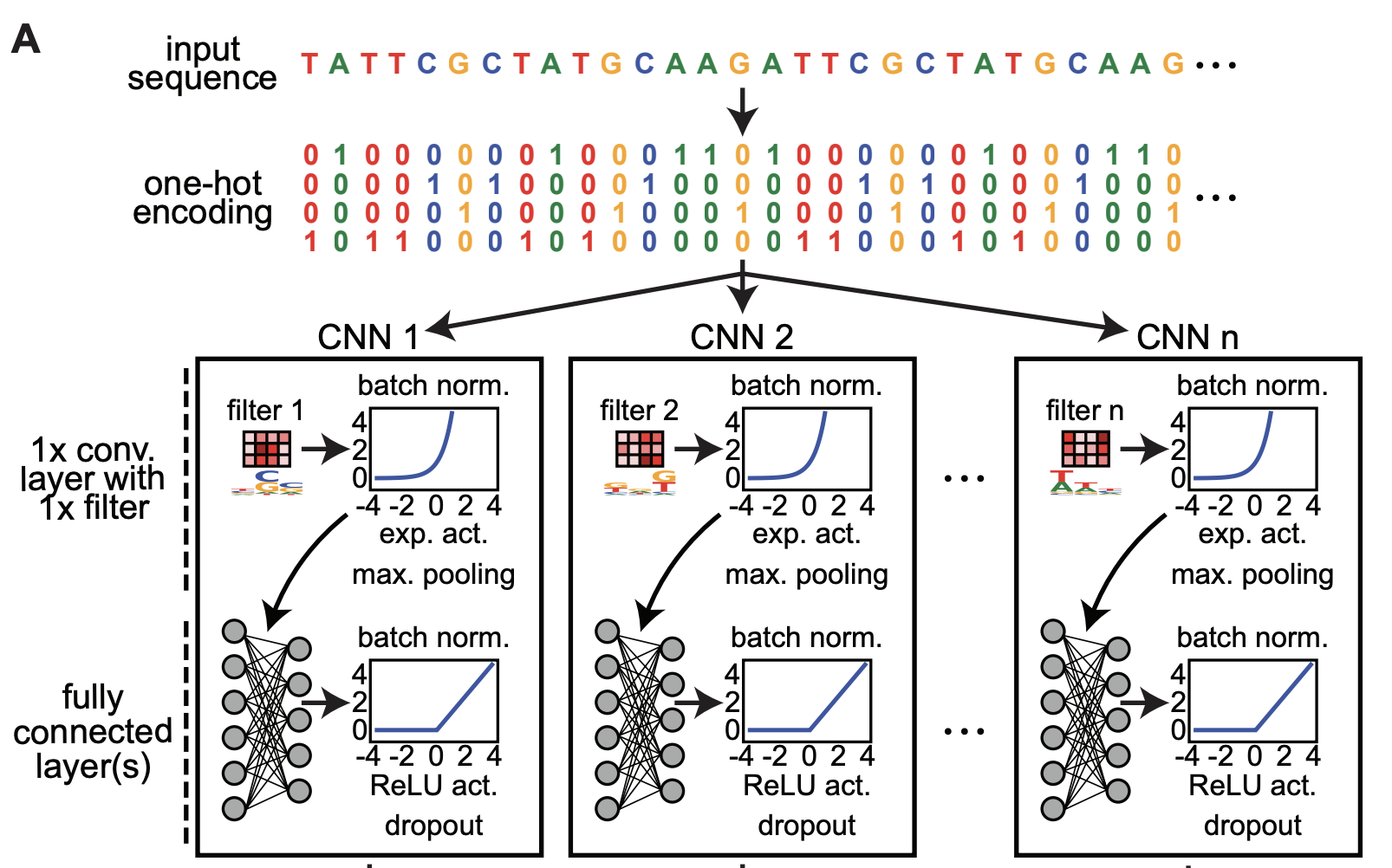
ExplaiNN: interpretable and transparent neural networks for genomics
Gherman Novakovsky, Oriol Fornes, Manu Saraswat, Sara Mostafavi, Wyeth W Wasserman. Genome Biology
2023
In silico discovery of small molecules for efficient stem cell differentiation into definitive endoderm
Gherman Novakovsky, Shugo Sasaki, ...Sara Mostafavi, Francis Lynn, Wyeth Wasserman. Stem Cell Reports.
2023
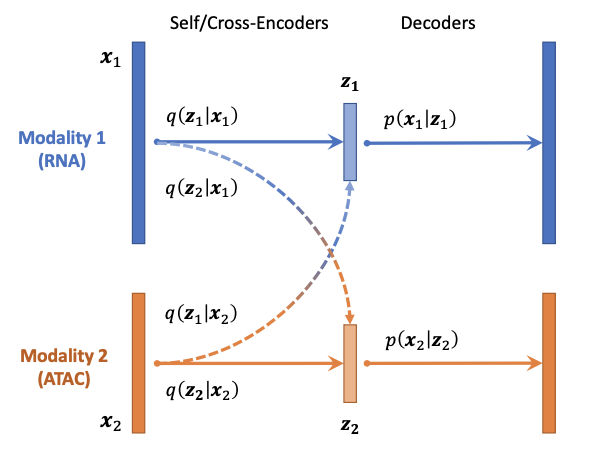
Cross-Linked Unified Embedding for cross-modality representation learning
Xinming Tu*, Zhi-Jie Cao*, Chen-Rui Xia, Sara Mostafavi$, Ge Gao$. NeurIPS (Oral).
2022
Obtaining genetics insights from deep learning via explainable artificial intelligence
Gherman Novakovsky*, Nick Dexter*, Maxwell Libbrecht$, Wyeth Wasserman$, Sara Mostafavi$. Nature Reviews Genetics.
2022
Mosaic loss of chromosome Y in aged human microglia
Michael Vermeulen, Richard Pearse, Tracy Young-Pearse, Sara Mostafavi. Genome Research. Cover article, Oct 2022 issue.
2022
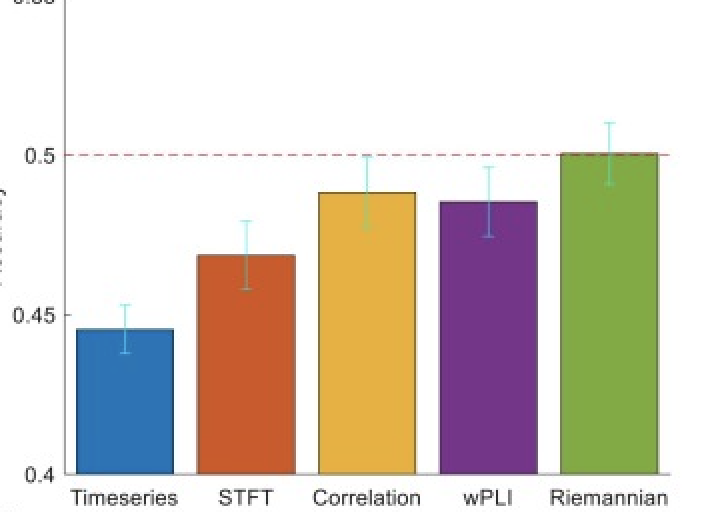
A practical guide to applying machine learning to infant EEG data
Bernard Ng*, Rebecca Reh, Sara Mostafavi$. Developmental Cognitive Neuroscience.
2022
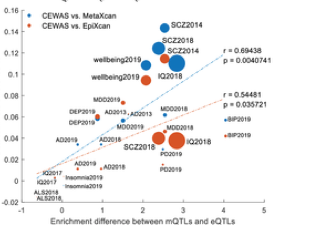
CEWAS: Cascading Epigenomic analysis for identifying disease genes from the regulatory landscape of GWAS variants
Bernard Ng, William Casazza, ..., Sara Mostafavi. PLOS Genetics.
2021
Biologically relevant transfer learning improves transcription factor binding prediction
Gherman Novakovsky, Manu Saraswat, Oriol Fornes, Sara Mostafavi, Wyeth Wasserman. Genome Biology.
2021
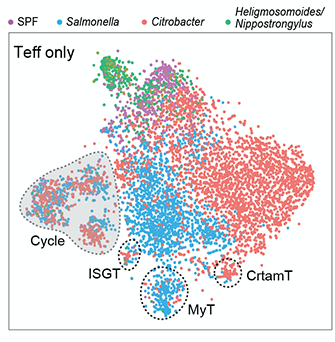
Gut CD4+ T cell phenotypes are a continuum molded by microbes, not by TH archetypes
Evgeny Kiner, Elijah Willie, Brinda Vijaykumar, ..,Sara Mostafavi, Diane Mathis, Christophe Benoist. Nature Immunology.
2021
Unified AI framework to uncover deep interrelationships between gene expression and Alzheimer's disease neuropathologies
Nicasia Beebe-Wang, Safiye Celik, ..., Sara Mostafavi$, Su-In Lee$. Nature Communications.
2021
Deep learning decodes the principles of differential gene expression
Shinya Tasaki, Chris Gaiteri, Sara Mostafavi, Yanling Wang. Nature Machine Intelligence.
2020
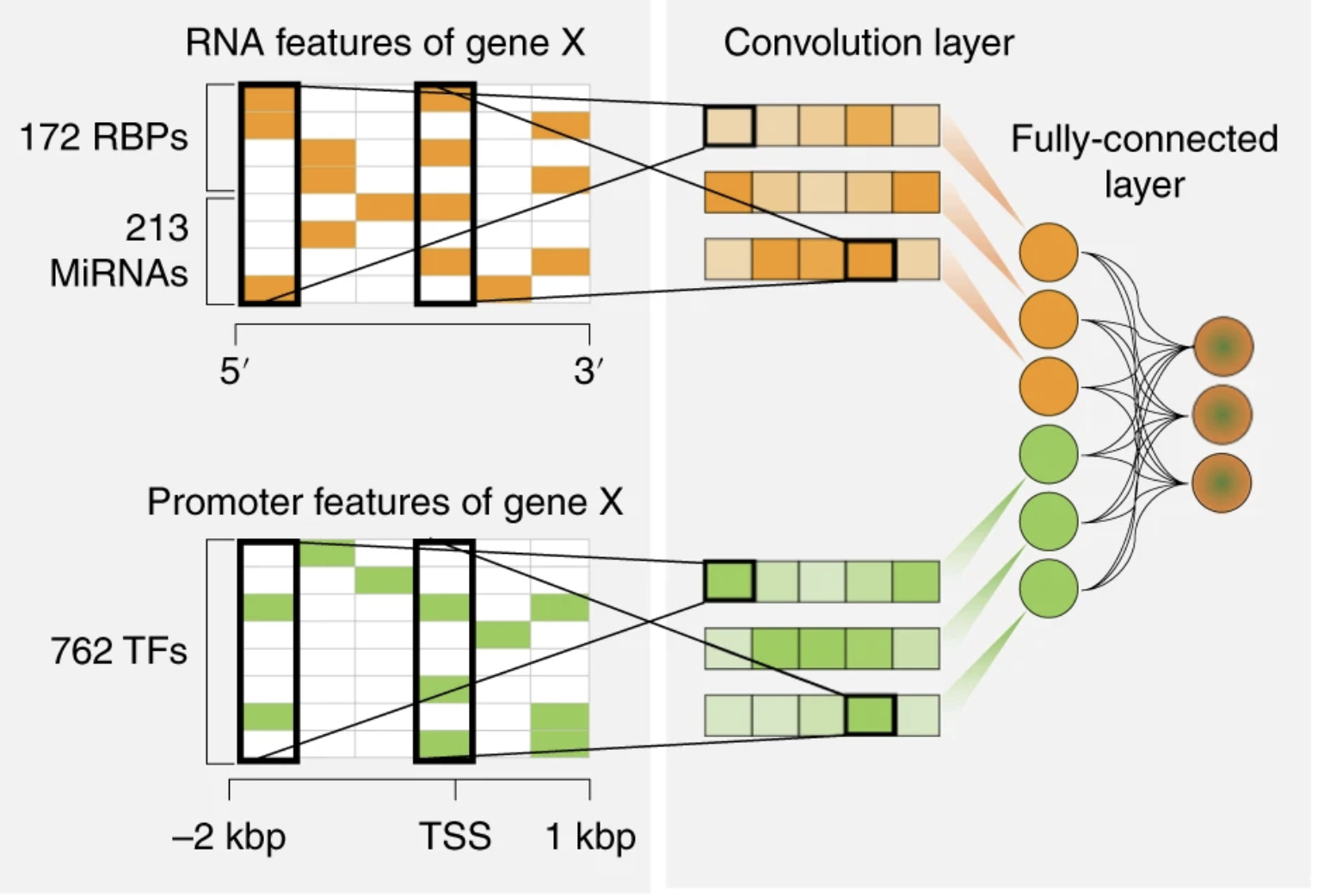
Deep learning of immune cell differentiation
Alexandra (Sasha) Maslova* , Ricardo Ramirez*, ... , Christophe Benoist$, Sara Mostafavi$. PNAS.
2020
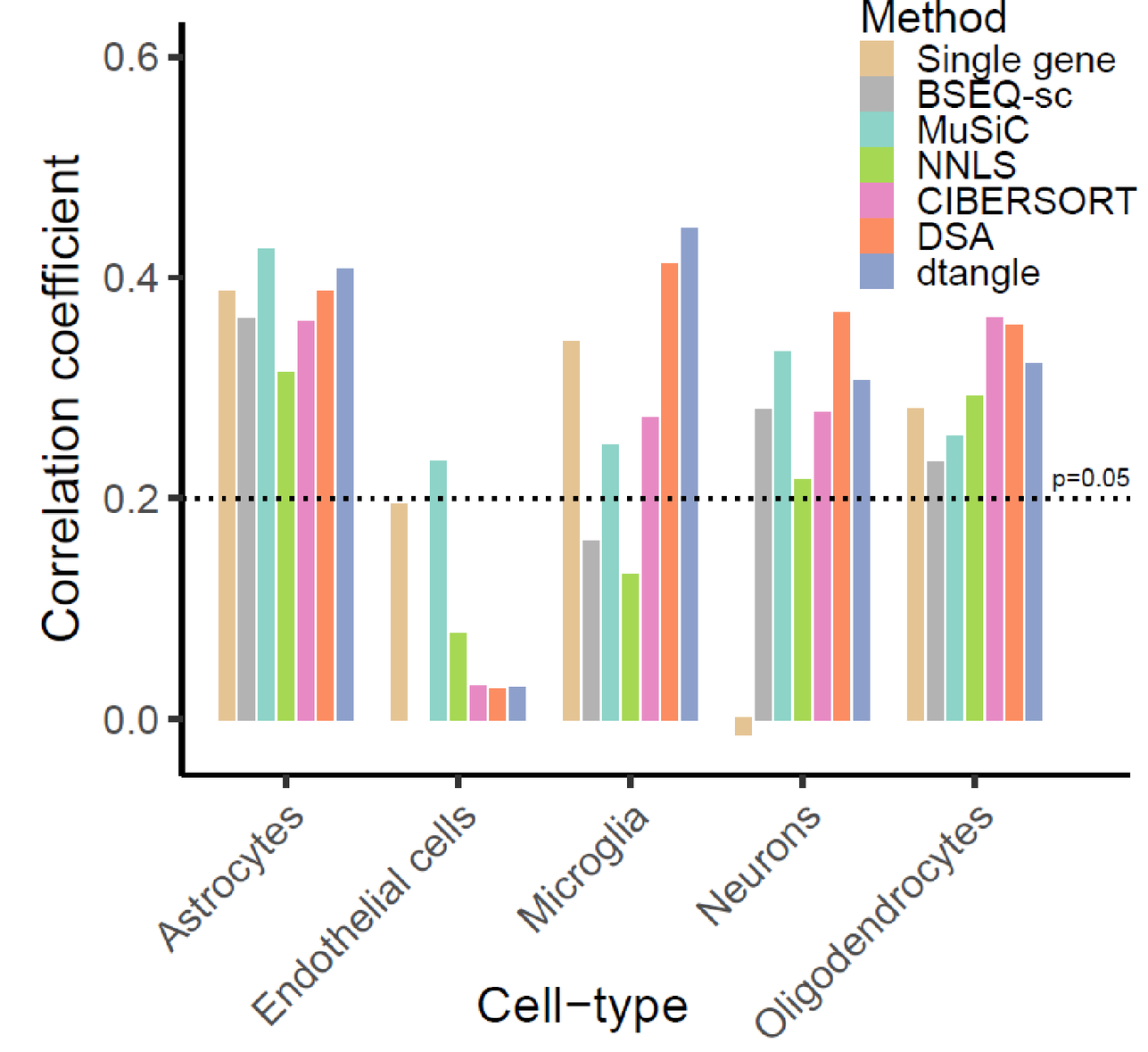
Deconvolving the contributions of cell-type heterogeneity on cortical gene expression
Ellis Patrick, Mariko Taga, Ayla Ergun, ... , Elizabeth Bradshaw$, Sara Mostafavi$. PLOS Computational Biology.
2020
metPropagate: network-guided propagation of metabolomic information for prioritization of metabolic disease genes.
Emma Graham Linck, Philip A. Richmond, ... , Mostafavi S. npj Genomic Medicine.
2020
Using Transcriptomic Hidden Variables to Infer Context-Specific Genotype Effects in the Brain.
Bernard Ng, William Casazza, ... , Mostafavi S. The American Journal of Human Genetics.
2019
Somatic mosaicism of sex chromosomes in the blood and brain.
Emma Graham, Mike Vermeulen, Badri Vardarjan, ... , Tracy Young-Pearse$,Mostafavi S$. Brain Research.
2019
The cis-Regulatory Atlas of the Mouse Immune System.
Yoshida H, Lereau C, ..., Mostafavi S$, Buenrostro J$, Benoist C$. Cell.
2019
A molecular network of the aging human brain provides insights into the pathology and cognitive decline of Alzheimer’s disease
Mostafavi S*, Gaiteri C*, ..., Bennett D, De Jager P. Nature Neuroscience.
2018
An xQTL map integrates the genetic architecture of the human brain's transcriptome and epigenome.
Ng B, White C, Klein HU, ..., Mostafavi S$, De Jager P$. Nature Neuroscience.
2017
Allele-specific expression reveals interactions between genetic variation and environment
Knowles D,Davis JR, Edgington H, ..., Mostafavi S, Montgomery SB, Battle AJ. Nature Methods.
2017
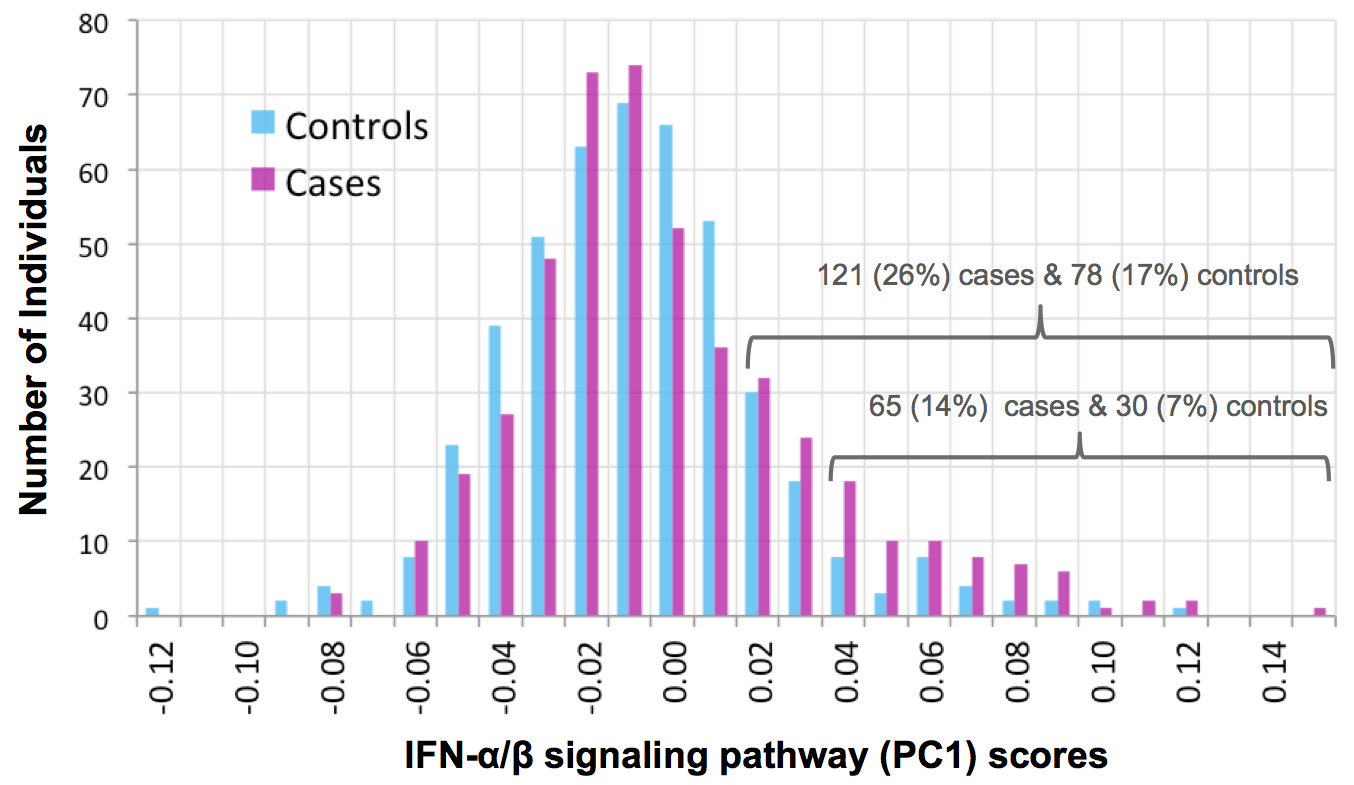
Parsing the interferon transcriptional network and its disease associations
Mostafavi S*, Yoshida H*, Moodley D, ..., Mathis D, Benoist C, and the ImmGen Consortium. Cell .
2016
Sharing and Specificity of Co-expression Networks across 35 Human Tissues
Pierson E, the GTEx Consortium, Koller D, Battle A$, Mostafavi S$. PLOS Computational Biology .
2015
Polarization of the effects of autoimmune and neurodegenerative risk alleles in leukocytes
Raj T, Rothamel K, Mostafavi S,..., De Jager PL. Science .
2014
Characterizing the genetic basis of transcriptome diversity through RNA-sequencing of 922 individuals
Battle A, Mostafavi S, Zhu X,..., Koller D. Genome Research .
2014
Normalizing RNA-sequencing data by modeling hidden covariates with prior knowledge
Mostafavi S, Battle A, Zhu X,..., Koller D. PLOS ONE .
2013
Type I interferon signaling genes in recurrent major depression: increased expression detected by whole-blood RNA sequencing
Mostafavi S, Battle A, Zhu X,..., Levinson DF. Molecular Psychiatry.
2013
Using prior biological knowledge when constructing gene regulatory networks
Mostafavi S, and Koller, D. ICML workshop on Machine Learning in Genomics and Genetics .
2012
Labeling nodes using three degrees of propagation
Mostafavi S, Goldenberg A, Morris Q. PLOS ONE .
2012
Predicting node characteristics from molecular networks
Mostafavi S, Goldenberg A, Morris Q. Methods in Molecular Biology .
2011
Fast integration of heterogeneous data sources for predicting gene function with limited annotation.
Mostafavi S and Morris Q. Bioinformatics .
2010
Using the Gene Ontology hierarchy when predicting gene function
Mostafavi S and Morris Q. Uncertainty in Artificial Intelligence (UAI) .
2009
GeneMANIA: a real-time multiple association network integration algorithm for predicting gene function
Mostafavi S, Ray D, Warde-Farley D, Grouios C, Morris Q. Genome Biology .
2008
Student Theses
Advancing our understanding of genome regulation via optimization of stem cell differentiation and interpretable deep learning
Gherman Novakovsky.
2022
Robust methods for identifying cluster structure in single-cell RNA-sequecing data
Elijah Willie.
2020
Improving the estimation of Y chromosome loss using high-throughput sequencing
Michael Vermeulen.
2020
Understanding gene regulatory mechanisms of mouse immune cells using a convolutional neural network
Alexandra (Sasha) Maslova.
2020
Integration of genomic and metabolomic data for the prioritization of rare disease variants.
Emma Graham.
2018
Molecular interpretation of genome-wide association studies using multiomics analysis.
Farnush Farhadi Hassan Kiadeh.
2018
Estimating cell type proportions in human cord blood samples from DNAm arrays.
Louie Dinh.
2017